Microsoft Patents an AI Pipeline That Generates New Molecules From Chemical Neighbors
Instead of asking an AI to invent molecules from scratch, Microsoft's new patent describes a smarter approach: find the chemical neighbors of a known compound first, then fine-tune a generative model on that richer neighborhood — nudging the AI toward territory that's already proven interesting.
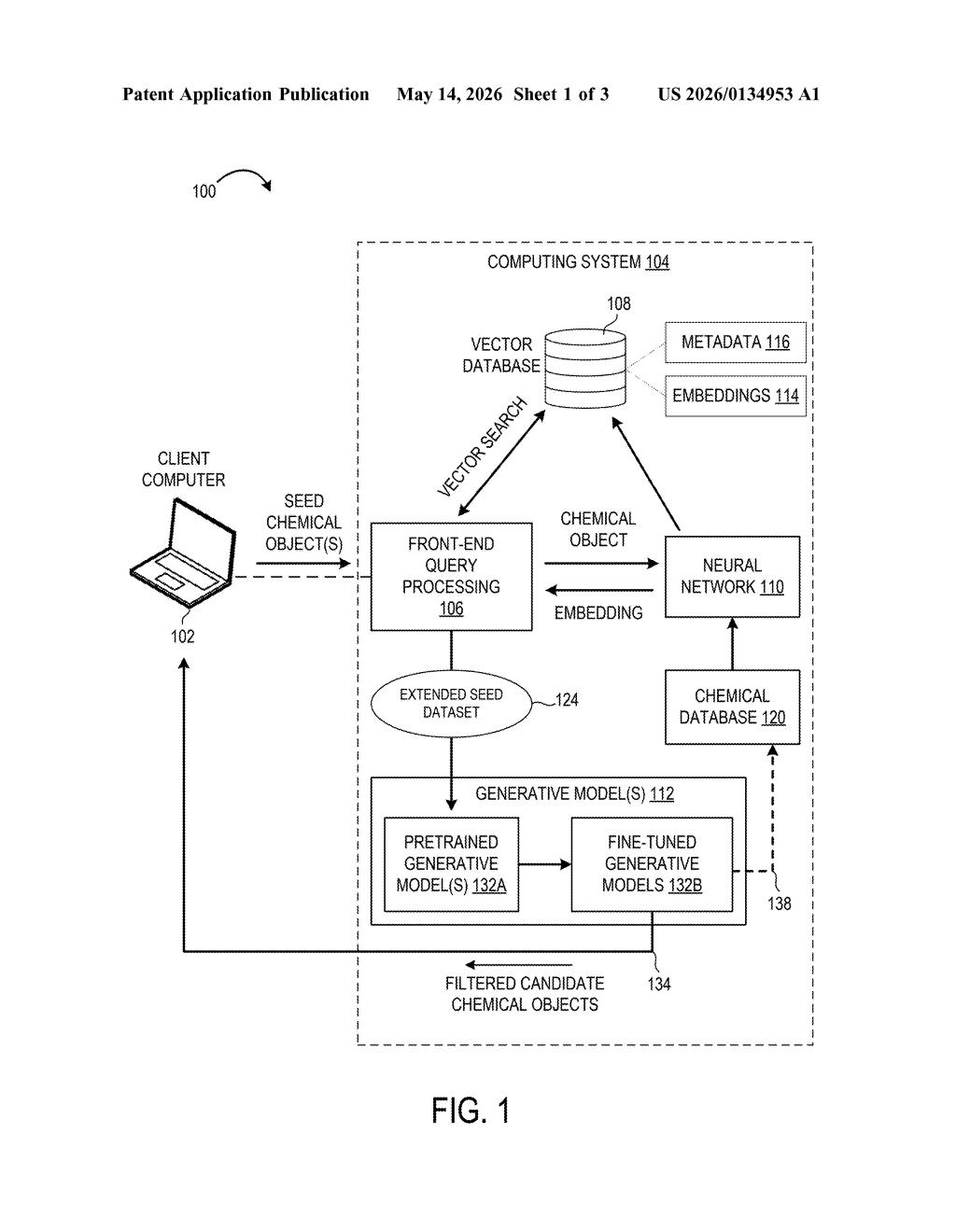
How Microsoft's system designs new molecules from examples
Imagine you're a chemist who discovered one promising drug compound and you want to find more molecules like it. You could ask an AI to generate random new ones, but most would be useless. Microsoft's patented approach is more like asking a very smart librarian to first pull every similar compound from a giant database, and then train a specialized AI on that curated shortlist before generating anything new.
Here's how it plays out for you as a researcher: you hand the system your starting molecule — the "seed." The system converts it into a mathematical fingerprint, scans a massive database for chemically similar fingerprints, and assembles an expanded training set. Then it takes a pre-existing generative AI model and fine-tunes it on just that expanded set.
The result is a generative model that's been narrowly focused on your chemical neighborhood of interest — meaning the molecules it proposes are more likely to share useful properties with your starting compound, rather than being random shots in the dark.
How the vector search steers the generative model
The patent describes a multi-stage pipeline built around a key insight: vector similarity search (finding items that are mathematically "close" in a high-dimensional space) can do the heavy lifting of chemical exploration before any new molecule is generated.
Here's the sequence the patent lays out:
- A researcher provides one or more seed chemical objects — molecules described in a machine-readable format like SMILES strings or molecular graphs.
- A neural network encoder converts each seed into an embedding (think of it as a dense numerical fingerprint that captures the molecule's chemical identity and structure).
- Those embeddings are compared against millions of pre-computed embeddings stored in a vector database — a specialized store optimized for fast nearest-neighbor lookups, similar to how image-search engines find visually similar photos.
- The search returns a set of chemically similar molecules, which are merged with the original seeds into an extended seed dataset.
- A pretrained generative model (one already trained broadly on large chemical datasets) is then fine-tuned on this focused dataset, producing a specialized model primed to generate molecules in the same chemical neighborhood.
The architecture separates two concerns cleanly: retrieval (what chemical space is relevant?) and generation (what new molecules should we propose?). That separation lets researchers swap in different encoders, databases, or base generative models without redesigning the whole pipeline.
What this means for AI-assisted drug discovery
Drug discovery lives and dies by chemical space navigation — the ability to explore millions of possible molecules efficiently. Fine-tuning a generative model on a similarity-expanded dataset is a meaningful technique because it sidesteps one of the core failure modes of generative AI in chemistry: producing molecules that are synthetically implausible or structurally irrelevant to the target of interest. By anchoring generation in a retrieved neighborhood, the system inherits the implicit chemical wisdom already encoded in the database.
For you as a researcher or biotech company, this could translate to faster lead optimization cycles — you start with a hit compound, and the system helps you explore analogs systematically rather than randomly. Microsoft is clearly positioning Azure AI infrastructure (vector databases, fine-tuning pipelines) as the backbone for life sciences workloads, and this patent is a concrete signal of that ambition.
This is a genuinely interesting application of retrieval-augmented generation (RAG) — a technique that's dominated NLP for the past two years — applied to molecular design. The core idea of using vector search to bootstrap fine-tuning data is clean and practically useful. It won't single-handedly transform pharma, but it's exactly the kind of infrastructure patent that quietly becomes a standard building block in AI-assisted drug discovery platforms.
Get one Big Tech patent every Sunday
Plain English, intelligent commentary, no hype. Free.
Editorial commentary on a publicly published patent application. Not legal advice.
