Microsoft Patents a Three-Phase AI System for Predicting Protein Shape Ensembles
Most protein-folding tools give you one shape. Microsoft's new patent targets the harder problem: a protein isn't a single rigid sculpture — it's a crowd of wiggling conformations, and predicting that whole crowd is what drug designers actually need.
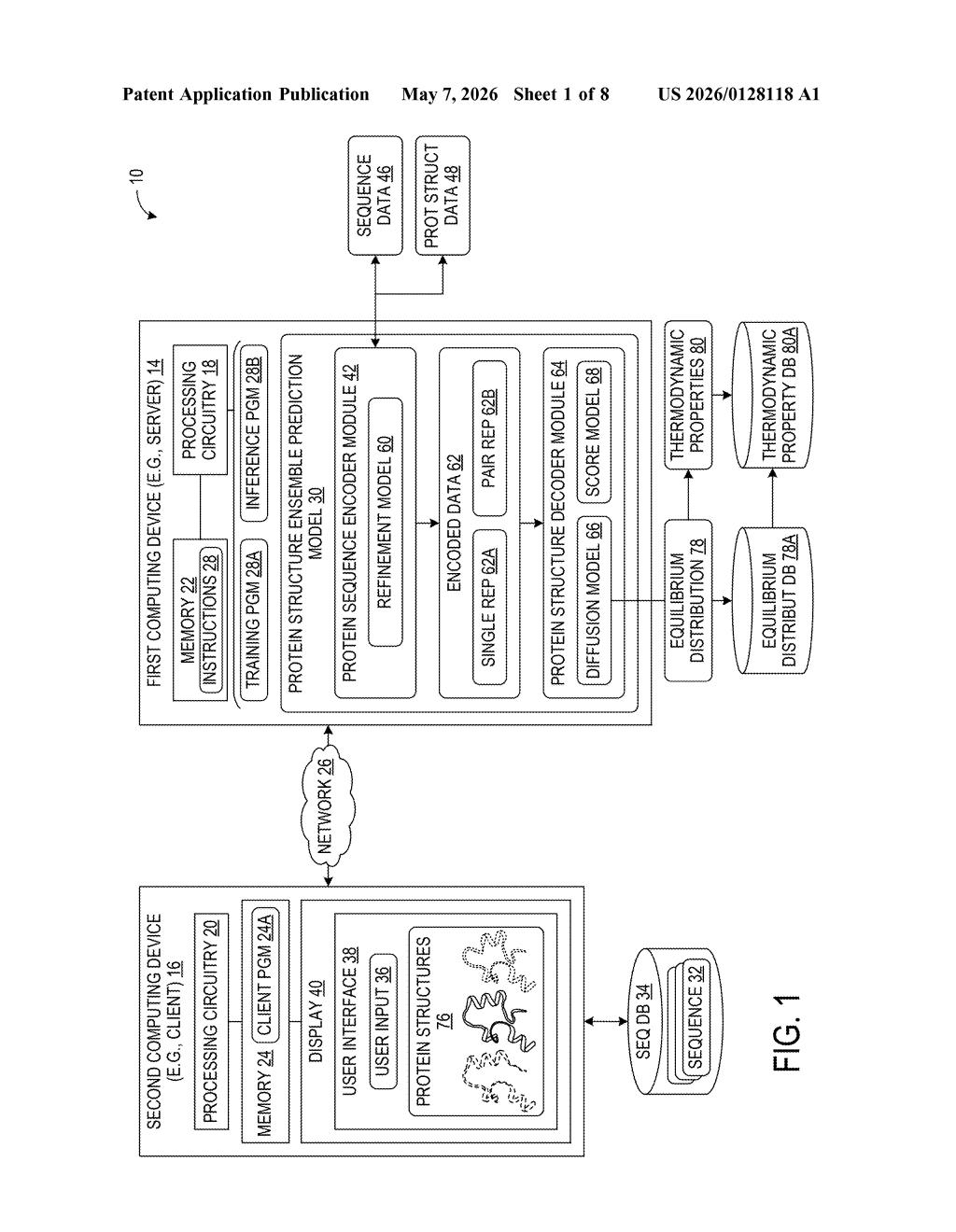
What Microsoft's protein ensemble predictor actually does
Imagine trying to design a key for a lock, but the lock keeps changing shape slightly depending on temperature, nearby molecules, or random thermal jostling. That's the real challenge in drug discovery — a protein target doesn't hold still, and a drug that only fits one snapshot of it may fail in the body.
Microsoft's patent describes an AI system trained to predict not just one protein structure, but a whole ensemble — a realistic spread of shapes that protein might adopt. Think of it like asking a weather model for a probability distribution of tomorrow's temperature rather than a single number.
The system learns in three stages: first from a large synthetic dataset, then by comparing its guesses against physics-based simulations, and finally by tuning itself so its predicted spread of shapes matches real thermodynamic properties. The result is a model that can generate plausible protein conformations in fifteen steps or fewer — fast enough to be practical.
How the three-phase diffusion training pipeline works
The patent describes a protein structure ensemble prediction model — a diffusion model (the same class of AI behind image generators like DALL-E, but applied to 3D molecular coordinates) that learns to output a distribution of plausible protein shapes rather than a single structure.
Training happens in three distinct phases:
- Phase 1 — Synthetic bootstrapping: The model ingests a large synthetic dataset of protein sequences, identifies sequences whose predicted structures are highly variable (structurally heterogeneous), clusters those structures by shape similarity, prunes out disordered or single-member clusters, and trains the diffusion model on the resulting clean pairs.
- Phase 2 — Molecular dynamics grounding: The model is shown structures sampled from molecular dynamics (MD) simulations — physics-based computer simulations that model how atoms move over time. It corrupts a known structure, tries to reconstruct it, and is penalized for errors versus the MD ground truth. This anchors the model in physical reality.
- Phase 3 — Thermodynamic property alignment: The model samples multiple structures for a given sequence, predicts a thermodynamic property (such as free energy or population weights) of that ensemble, and is backpropagated (error-corrected) based on how far off that prediction is from real measured values.
Critically, the diffusion model is designed to converge in fifteen or fewer denoising steps — a meaningful efficiency target that makes ensemble sampling computationally tractable.
Why predicting multiple protein shapes matters for drug design
Single-structure protein prediction tools like AlphaFold transformed structural biology, but the field has been pushing hard toward ensemble prediction because biological function — and drug binding — depends on the full range of shapes a protein visits. A system that can cheaply and accurately sample that range could meaningfully accelerate hit identification and lead optimization in pharmaceutical pipelines.
For Microsoft, this filing signals a serious push into computational biology infrastructure, an area where Google DeepMind (AlphaFold), Meta (ESMFold), and several biotech startups are all competing. The three-phase training strategy — synthetic data, MD simulation grounding, thermodynamic property supervision — reads like a thoughtful attempt to combine data efficiency with physical correctness, which is the core tension in this space.
This is a genuinely interesting patent because it targets the right hard problem — ensemble prediction with thermodynamic fidelity — rather than incremental improvements on single-structure folding. The three-phase training pipeline is sophisticated and clearly designed by people who understand both the ML and the biophysics. Whether Microsoft can translate this into a competitive product against entrenched academic and commercial tools is the real question, but the technical ambition here is real.
Get one Big Tech patent every Sunday
Plain English, intelligent commentary, no hype. Free.
Editorial commentary on a publicly published patent application. Not legal advice. Patentlyze may earn a commission if you click an affiliate link and make a purchase. This doesn't affect what we cover or how we cover it.
